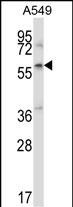
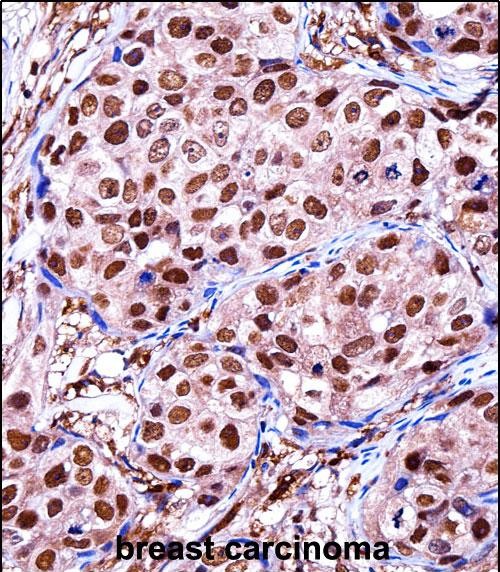
ZBTB7A Antibody (C-term)
Affinity Purified Rabbit Polyclonal Antibody (Pab)
- 产品详情
- 文献引用 : 1
- 实验流程
- 背景知识
Application
| WB, IHC-P, E |
|---|---|
| Primary Accession | O95365 |
| Other Accession | NP_056982.1 |
| Reactivity | Human |
| Host | Rabbit |
| Clonality | Polyclonal |
| Isotype | Rabbit IgG |
| Calculated MW | 61439 Da |
| Antigen Region | 514-542 aa |
| Gene ID | 51341 |
|---|---|
| Other Names | Zinc finger and BTB domain-containing protein 7A, Factor binding IST protein 1, FBI-1, Factor that binds to inducer of short transcripts protein 1, HIV-1 1st-binding protein 1, Leukemia/lymphoma-related factor, POZ and Krueppel erythroid myeloid ontogenic factor, POK erythroid myeloid ontogenic factor, Pokemon, TTF-I-interacting peptide 21, TIP21, Zinc finger protein 857A, ZBTB7A, FBI1, LRF, ZBTB7, ZNF857A |
| Target/Specificity | This ZBTB7A antibody is generated from rabbits immunized with a KLH conjugated synthetic peptide between 514-542 amino acids from the C-terminal region of human ZBTB7A. |
| Dilution | WB~~1:1000 IHC-P~~1:100~500 E~~Use at an assay dependent concentration. |
| Format | Purified polyclonal antibody supplied in PBS with 0.09% (W/V) sodium azide. This antibody is purified through a protein A column, followed by peptide affinity purification. |
| Storage | Maintain refrigerated at 2-8°C for up to 2 weeks. For long term storage store at -20°C in small aliquots to prevent freeze-thaw cycles. |
| Precautions | ZBTB7A Antibody (C-term) is for research use only and not for use in diagnostic or therapeutic procedures. |
| Name | ZBTB7A (HGNC:18078) |
|---|---|
| Function | Transcription factor that represses the transcription of a wide range of genes involved in cell proliferation and differentiation (PubMed:14701838, PubMed:17595526, PubMed:20812024, PubMed:25514493, PubMed:26455326, PubMed:26816381). Directly and specifically binds to the consensus sequence 5'-[GA][CA]GACCCCCCCCC-3' and represses transcription both by regulating the organization of chromatin and through the direct recruitment of transcription factors to gene regulatory regions (PubMed:12004059, PubMed:17595526, PubMed:20812024, PubMed:25514493, PubMed:26816381). Negatively regulates SMAD4 transcriptional activity in the TGF-beta signaling pathway through these two mechanisms (PubMed:25514493). That is, recruits the chromatin regulator HDAC1 to the SMAD4-DNA complex and in parallel prevents the recruitment of the transcriptional activators CREBBP and EP300 (PubMed:25514493). Collaborates with transcription factors like RELA to modify the accessibility of gene transcription regulatory regions to secondary transcription factors (By similarity). Also directly interacts with transcription factors like SP1 to prevent their binding to DNA (PubMed:12004059). Functions as an androgen receptor/AR transcriptional corepressor by recruiting NCOR1 and NCOR2 to the androgen response elements/ARE on target genes (PubMed:20812024). Thereby, negatively regulates androgen receptor signaling and androgen- induced cell proliferation (PubMed:20812024). Involved in the switch between fetal and adult globin expression during erythroid cells maturation (PubMed:26816381). Through its interaction with the NuRD complex regulates chromatin at the fetal globin genes to repress their transcription (PubMed:26816381). Specifically represses the transcription of the tumor suppressor ARF isoform from the CDKN2A gene (By similarity). Efficiently abrogates E2F1-dependent CDKN2A transactivation (By similarity). Regulates chondrogenesis through the transcriptional repression of specific genes via a mechanism that also requires histone deacetylation (By similarity). Regulates cell proliferation through the transcriptional regulation of genes involved in glycolysis (PubMed:26455326). Involved in adipogenesis through the regulation of genes involved in adipocyte differentiation (PubMed:14701838). Plays a key role in the differentiation of lymphoid progenitors into B and T lineages (By similarity). Promotes differentiation towards the B lineage by inhibiting the T-cell instructive Notch signaling pathway through the specific transcriptional repression of Notch downstream target genes (By similarity). Also regulates osteoclast differentiation (By similarity). May also play a role, independently of its transcriptional activity, in double-strand break repair via classical non-homologous end joining/cNHEJ (By similarity). Recruited to double-strand break sites on damage DNA, interacts with the DNA-dependent protein kinase complex and directly regulates its stability and activity in DNA repair (By similarity). May also modulate the splicing activity of KHDRBS1 toward BCL2L1 in a mechanism which is histone deacetylase-dependent and thereby negatively regulates the pro-apoptotic effect of KHDRBS1 (PubMed:24514149). |
| Cellular Location | Nucleus. Note=Recruited to double-strand break sites of damaged DNA. {ECO:0000250|UniProtKB:O88939} |
| Tissue Location | Widely expressed (PubMed:9927193). In normal thymus, expressed in medullary epithelial cells and Hassle's corpuscles (at protein level) (PubMed:15662416). In tonsil, expressed in squamous epithelium and germinal center lymphocytes (at protein level) (PubMed:15662416). Up-regulated in a subset of lymphomas, as well as in a subset of breast, lung, colon, prostate and bladder carcinomas (at protein level) (PubMed:15662416). Expressed in adipose tissues (PubMed:14701838). |
For Research Use Only. Not For Use In Diagnostic Procedures.

Provided below are standard protocols that you may find useful for product applications.
BACKGROUND
ZBTB7A plays a key role in the instruction of early lymphoid progenitors to develop into B lineage by repressing T-cell instructive Notch signals (By similarity). Specifically represses the transcription of the CDKN2A gene. Efficiently abrogates E2F1-dependent CDKN2A transactivation/de-repression. Binds to the consensus sequence 5'-[GA][CA]GACCCCCCCCC-3' (By similarity).
REFERENCES
Qu, H., et al. Cancer Invest. 28(6):672-678(2010)
Zu, X., et al. Cell. Mol. Biol. Lett. 15(2):260-271(2010)
Cui, J., et al. Mol. Biol. Rep. 37(3):1627-1632(2010)
Choi, W.I., et al. J. Biol. Chem. 284(19):12633-12644(2009)
Choi, W.I., et al. Cell. Physiol. Biochem. 23 (4-6), 359-370 (2009) :
终于等到您。ABCEPTA(百远生物)抗体产品。
点击下方“我要评价 ”按钮提交您的反馈信息,您的反馈和评价是我们最宝贵的财富之一,
我们将在1-3个工作日内处理您的反馈信息。
如有疑问,联系:0512-88856768 tech-china@abcepta.com.






















 癌症的基本特征包括细胞增殖、血管生成、迁移、凋亡逃避机制和细胞永生等。找到癌症发生过程中这些通路的关键标记物和对应的抗体用于检测至关重要。
癌症的基本特征包括细胞增殖、血管生成、迁移、凋亡逃避机制和细胞永生等。找到癌症发生过程中这些通路的关键标记物和对应的抗体用于检测至关重要。 为您推荐一个泛素化位点预测神器——泛素化分析工具,可以为您的蛋白的泛素化位点作出预测和评分。
为您推荐一个泛素化位点预测神器——泛素化分析工具,可以为您的蛋白的泛素化位点作出预测和评分。 细胞自噬受体图形绘图工具为你的蛋白的细胞受体结合位点作出预测和评分,识别结合到自噬通路中的蛋白是非常重要的,便于让我们理解自噬在正常生理、病理过程中的作用,如发育、细胞分化、神经退化性疾病、压力条件下、感染和癌症。
细胞自噬受体图形绘图工具为你的蛋白的细胞受体结合位点作出预测和评分,识别结合到自噬通路中的蛋白是非常重要的,便于让我们理解自噬在正常生理、病理过程中的作用,如发育、细胞分化、神经退化性疾病、压力条件下、感染和癌症。